Odor Management at Landfills: Part 2 - A Novel Biosensor for Measuring Odors at Landfills
November 11, 2021

By Daniel Meeroff, Ph.D. and Sharmily Rahman
The absence of an objective measurement technique for nuisance odor strength creates difficulties for regulatory agencies to draft effective rules for solid waste managers to deal with odor complaints.
To address this problem, researchers at Florida Atlantic University’s Laboratories for Engineered Environmental Solutions (Lab.EES) are working on a unique idea developed by Dr. Daniel E. Meeroff. The concept takes inspiration from human biology and how the sense of smell actually works to design a biosensor that can objectively quantify odors using copies of the human odorant binding protein (hOBPIIa).
These are the same proteins that shepherd airborne odorants to the olfactory receptors in the human nose, so that a signal can be sent to the brain to trigger a response to ambient odors.
There has been a recent surge in discovery of unique applications of odorant binding proteins (OBPs). For instance, OBPs isolated from pigs were used to trap the smell of cigarettes (Silva 2014).
Another line of research explored whether pig OBPs can make perfume last longer when sprayed on clothes. The use of advanced nano-biosensors derived from OBP isolated from cows had also been investigated to detect the presence of explosive and hazardous compounds in luggage storage facilities, airports and other public places (Ramoni et al. 2007). However, none of these applications used OBPs isolated from humans (Figure 1), and none for the purpose of measuring odorant concentrations.
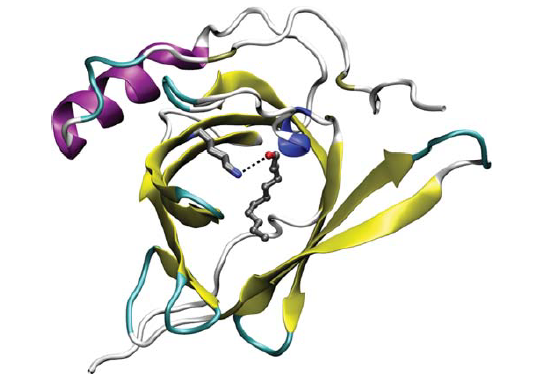
Figure 1: Tertiary structure of human odorant binding protein-2A (hOBPIIa) bonded with an aldehyde known as undecanal (in the middle) (Heydel et al. 2013)
Recent advances in biotechnology (Silva 2014) have allowed these highly specialized proteins (hOBPIIa) to be easily replicated in the lab to form the basis of an odorant-detecting biosensor. These proteins bind with a wide range of odorants in the micromolar range. So by adding a biomolecular fluorescent marker, which in this case is the fluorophore 1-AMA (Figure 2), binding is quantified using a common spectrofluorometer.
Since the human nervous system processes odor intensity based on the number of bound receptor sites, and the biosensor protein fluorescent response is concentration-dependent, this new technology will allow for authentic quantification of odors without subjective interpretation.
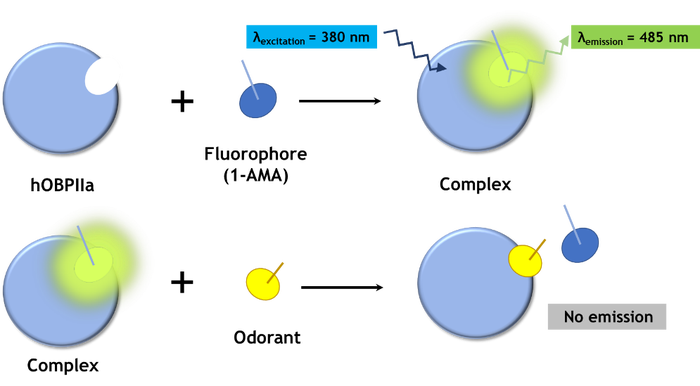
Figure 2: Mechanism of quantifying odorant gases with hOPBIIa by means of spectrofluorometric analysis
The analysis is conducted in a closed chamber containing the fluorescently tagged protein biosensor suspended in a phosphate buffer solution. Then air containing odorants is pumped into the chamber, and samples are collected periodically to measure the fluorescence intensity using a spectrofluorometer.
With time, the peak fluorescent intensity decreases as more of the protein releases the bond with the fluorophore to bind with the odorants. This provides an indirect measure of how much odorant is present in the solution.
This decreasing trend of fluorescence intensity proceeds until all proteins in the chamber are saturated with odorants. Figure 3 represents a general decreasing pattern of the peak fluorescence intensity against increasing odorant gas concentration in the biosensor solution attained from the experiments. The linear portion of the curve is the quantitation range.
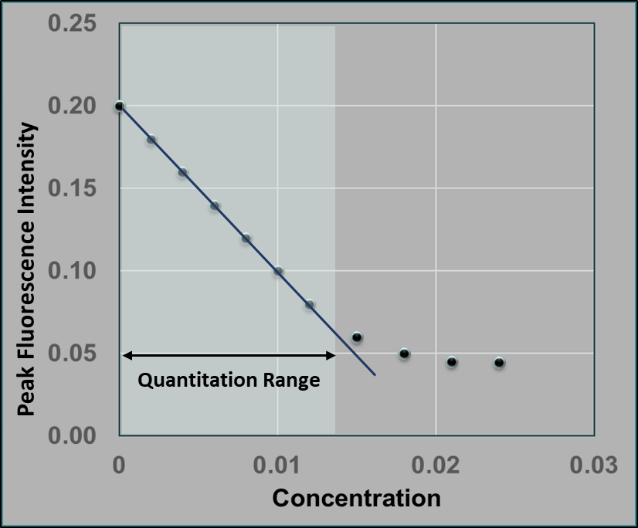
Figure 3: The decreasing trend of peak emission intensity against odorant gas concentrations in the biosensor solution up to the quantitation range
A number of common odorants found in landfills and other industrial facilities have already been tested in the laboratory using the new biosensor, and the results thus far have been really encouraging.
Tests have been conducted using common odorants such as hydrogen sulfide, ammonia, methyl mercaptan, and methane, as well as mixtures (Mixture 1: hydrogen sulfide, methane and carbon monoxide; Mixture 2: ammonia and methane), all of which are typically present in landfills. Each odorant gas and mixtures of gases demonstrated a unique pattern of decreasing fluorescence, which can be used in identifying the type of odor reacting with the biosensor solution.
Among all the odorant gases tested, ammonia is found to saturate the biosensor solution the fastest due to its higher solubility. For each of the experiments, around 180 μg of protein was used and was found to detect approximately 35-45 μg of hydrogen sulfide, 10-20 μg of ammonia, 80-95 μg of methyl mercaptan, 15 μg of methane depending on the pumping rate.
Table 1 shows the quantitation ranges and masses of the pure odorant gases detected by the biosensor solution for different flow rates (measured in cubic feet per second) to the quantitation range, which is the analytical response time (on the order of seconds).
Table 1: Quantitation ranges and masses of the pure odorant gases tested at different flow rates
Odorant gas | Quantitation Range (seconds) | Mass Range (μg) |
Q = 0.018 cfm | Q = 0.025 cfm | Q = 0.032 cfm |
Hydrogen Sulfide | 120 | 110 |
Ammonia | 90 | 75 |
Methyl Mercaptan | 95 | 75 |
Methane | 100 | - |
For the mixtures, since there is more than one gas present, the total amount of odor is determined in a competitive assay. Currently, FAU researchers are attempting to work out which gas molecules actually bind with the protein and if there is a preferential affinity for certain gases.
While the results have been very encouraging, there is still more work to do before the technology becomes commercially available. Thus far, experiments have only been conducted in a laboratory setting. The ultimate goal is to use develop a hand-held device with a built-in pump, integrated flow-through spectrofluorometer and reactor chamber cartridge.
All of these components are commercially available and relatively inexpensive to deploy at this scale. Further work needs to be done to determine if the biosensor can be regenerated after a single use so that the cartridge can be reused many times. Purging with nitrogen gas or zero air has been tested for 1-2 times the quantitation range, but initial experiments indicate that full regeneration may take longer.
To effectively deal with nuisance odors, an objective measurement technique that can quantify odors rapidly in the field is needed.
Since OBPs can be readily synthesized from recombinant DNA in bacteria, the idea of using these proteins as biosensors to quantify odors is a step forward in achieving this goal at low cost. FAU has managed to miniaturize the chamber to reduce losses, lower the response time, and reduce cost of manufacturing proteins compared to previous work.
This new rapid biosensor technology will allow objective measurements of odor concentrations in real-time, potentially transforming the way that nuisance odors are regulated and monitored, not only for solid waste managers but also for other industries as well. Dealing with odor complaints is an important part of the public image of solid waste facilities.
Armed with a revolutionary new technology, the industry will be better able to respond to odor issues with improved certainty and regulatory clarity, saving money and potentially repairing the sometimes confrontational relationship with their nearest neighbors.
###
This is part two in a two-part series. Read Part One here.
References:
Heydel, J. M., Coelho, A., Thiebaud, N., Legendre, A., Bon, A. M. L., Faure, P., ... & Briand, L. (2013). Odorant‐binding proteins and xenobiotic metabolizing enzymes: implications in olfactory perireceptor events. The Anatomical Record, 296(9), 1333-1345. https://doi.org/10.1002/ar.22735
Ramoni, R., Bellucci, S., Grycznyski, I., Grycznyski, Z., Grolli, S., Staiano, M., ... & Conti, V. (2007). The protein scaffold of the lipocalin odorant-binding protein is suitable for the design of new biosensors for the detection of explosive components. Journal of Physics: Condensed Matter, 19(39), 395012.
Silva, C., Matamá, T., Azoia, N. G., Mansilha, C., Casal, M., & Cavaco-Paulo, A. (2014). Odorant binding proteins: a biotechnological tool for odour control. Applied microbiology and biotechnology, 98(8), 3629-3638.
Vidovic, M. (2017). Meteorological Conditions Affecting the Dispersion of Landfill Odor Complaints. Florida Atlantic University.
*******
Authors:
Daniel Meeroff, Ph.D.
Dr. Meeroff is Professor and Associate Chair of the Department of Civil, Environmental & Geomatics Engineering at Florida Atlantic University. He is also the Director of the Laboratories for Engineered Environmental Solutions (labees.civil.fau.edu) with over 20 years of experience in solid/hazardous waste management, pollution prevention, and environmental engineering research. Email:[email protected]
Sharmily Rahman
Mrs. Rahman is a doctoral candidate in the Department of Civil, Environmental & Geomatics Engineering at Florida Atlantic University. She was team leader for FAU’s first place winning entry in the 2019 SWANA International Student Design Competition. Email:[email protected]
You May Also Like


